Single-cell RNA-seq of childhood medulloblastoma
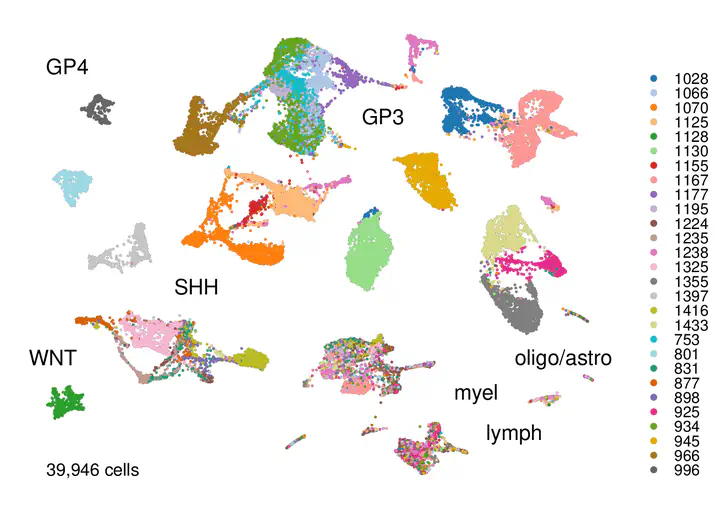
We explored cellular heterogeneity in medulloblastoma (MB) using single-cell RNA sequencing (scRNA-seq), immunohistochemistry and deconvolution of bulk transcriptomic data. Over 45,000 cells from 28 patients from all main subgroups of medulloblastoma (1 WNT, 9 SHH, 7 GP3 and 11 GP4) were clustered using Harmony alignment to identify conserved subpopulations.
Accompanies the manuscript “Neoplastic and immune single cell transcriptomics define subgroup-specific intra-tumoral heterogeneity of childhood medulloblastoma”
Data and code availability
scRNA-seq and methylation data have been deposited in the National Center for Biotechnology Information Gene Expression Omnibus (GEO) database. GSE156053
A UCSC cellbrowser is available for interactive exploration of the data.
Analysis scripts (Rmarkdown) are hosted on github: rnabioco/medulloblast
Supported by
National Institutes of Health (R01 CA237608-01)
Cancer League of Colorado (research grant AWD 173796-SV)
Tanner Seebaum Foundation
Morgan Adams Foundation
University of Colorado’s NIH/NCI Cancer Center (P30CA046934)